|
4/6/2023 0 Comments Geneious prime tutorial
In this protocol we discuss and outline the process of de novo assembly for small to medium sized genomes. Genome assembly refers to the process of taking a large number of short DNA sequences and putting them back together to create a representation of the original chromosomes from which the DNA originated. De novo genome assemblies assume no prior knowledge of the source DNA sequence length, layout or composition. In a genome sequencing project, the DNA of the target organism is broken up into millions of small pieces and read on a sequencing machine. These “reads” vary from 20 to 1000 nucleotide base pairs (bp) in length depending on the sequencing method used. Typically for Illumina type short read sequencing, reads of length 36 - 150 bp are produced. These reads can be either “single ended” as described above or “paired end.” A good summary of other types of DNA sequencing can be found here. Paired end reads are produced when the fragment size used in the sequencing process is much longer (typically 250 - 500 bp long) and the ends of the fragment are read in towards the middle. One from the left hand end of a fragment and one from the right with a known separation distance between them. (The known separation distance is actually a distribution with a mean and standard deviation as not all original fragments are of the same length.) This extra information contained in the paired end reads can be useful for helping to tie pieces of sequence together during the assembly process. The goal of a sequence assembler is to produce long contiguous pieces of sequence (contigs) from these reads. The contigs are sometimes then ordered and oriented in relation to one another to form scaffolds. “agents” and collaboration modes) - the best advice I can give is to download it, do the tutorial and test drive it on a bioinformatics task that you regularly conduct - it won't disappoint.The distances between pairs of a set of paired end reads is useful information for this purpose. 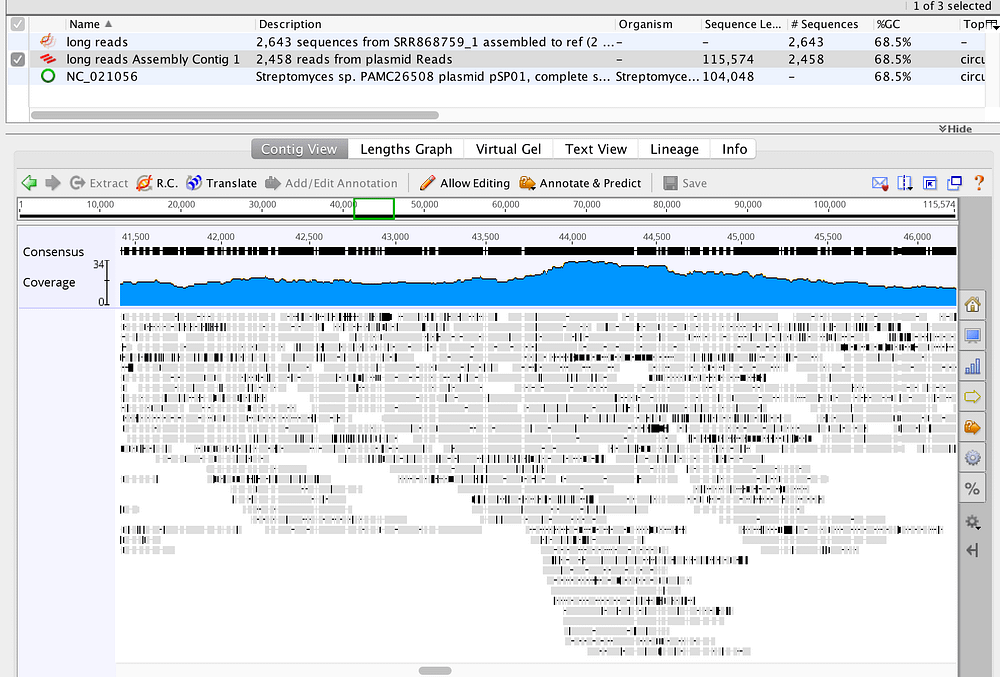
The software almost has too many features to comment on in any one review (e.g. Based on the hours of time this program has saved me already I would not even blink at the academic price tag of $133 (USD) - or cheaper for students. The Geneious team has made available a freeware version and a pro version that (for a price) allows you to access more functionality. Moreover, the software is structured in such a way that “new” plugins can be written and incorporated with relative ease. My request to provide a variety of nucleotide colour schemes was implemented within 2 weeks. As is the case with any new software there are a few features that don't work quite as well as you might like - but the good news is that if you write to them they will likely fix it up asap. With the integration of sequence chromatograph viewing and editing of DNA sequences/alignments (new in version 2) Geneious is now firmly embedded in my research practices. If you are looking for a piece of bioinformatics software for teaching purposes look no further than Geneious. Students did a Genbank search, downloaded files, did an alignment and built a phylogenetic tree (with bootstrapping) all in the blink of an eye. Earlier this year I used the Beta-release in some undergraduate teaching labs - with great success. While the software has some “bell and whistles” for more complex tasks one of its key attributes is a simple user interface. The user interface of Geneious is both attractive and intuitive. The software logo “research in a flash” accurately describe the potential of Geneious - only months after its release it has become the “Swiss army knife” of bioinformatics software. Current and future releases of Geneious (offer a solution to (most) of these frustrations. 
Chopping and changing between different file formats and at times different operating systems (Mac, PC, Unix/Linux) has been a frustrating and time-consuming exercise. 
For a long time scientists who generate or analyse DNA sequences have been using multiple pieces of software to: 1) view chromatographs, 2) trim sequences 3) Blast data 4) align sequences 5) build phylogenetic trees.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
